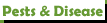according to Butterworth's cladogram, is Lophophora williamsii more related to Obegonia denegrii than to Lophophora diffusa, or am I reading this wrong?
Quite possibly. Other studies have also shown that Obregonia is simply a Lophophora with elongated tubercles. Note also the position of Acharagma. The exact relationships are not statistically certain in Butterworth's paper, but other work has shown that Acharagma is certainly closer to Lophophora than to anything else.
Echinocactus texensis is called Homalocephala texensis instead ,by the author, and for a good reason!
Again the probabilities are a little low for this relationships. Other studies place it with the other Echinocactus and separate from Astrophytum. They're all close though. Crozier, after splitting Coryphantha and Mammillaria into a thousand pieces, decides that Homocephala should stay lumped in Echinocactus. The traditional differences include the flattened growth habit, many sharp ribs (but see E. polycephalus with sharp ribs), pubescent spines (but again see E. polycephalus), and bright red fleshy fruit protruding from the wool.
We have witnessed all the above miss-classifications, why would Ferocactus grusonii sound wrong?
E. grusonii is only related to Ferocactus on the maternal line. It is left in the genus Echinocactus because that's what the flowers, fruit, and seed look like and they are considered more taxonomically significant than shiny skin. A genus Ferocactus including E. grusonii would be a disaster because E. grusonii has none of the characters that set Ferocactus apart from the other Echinocactus species. You want Ferocactus grusonii, feel free to write it up and have it published, but I doubt it will catch on. I think you're cherry-picking data (ie. maternal line chloroplast DNA) to support a pet theory when far more data contradicts it.
The numbers representing the relationship between E. ingens and E. horizonthalonius are 76/1. They aren't very strong affiliated either, are they?
These numbers don't mean that these species are not closely related, merely that this study does not strongly prove that relationship. Possibly they aren't close, possibly they are. Others studies show that they are certainlly related but not as close as, for example, E. parryi and E. polycephalus.
In Nyffeler's work we see E. platyacanthus related to Astrophytum myriostigma.
Where is this paper? I haven't been able to find it.
is Echinocactus, numbering now only six species, a well established and solid genus or a leftover from the past
It is certainly a diverse genus, very much a relic of the original very large genus. I don't think anyone would have created it as it stands, but you can't disband it without something else to put in it's place. Is it so hard to believe that Astrophytum capricorne would be an Echinocactus? Compare the flowers and fruits some time, they are remarkably similar, certainly more variable within each genus than they are different from eachother. The distinctive skin flecks of Astrophytum could easily have been ignored if the genus was being created today with all that we now know.
even the genus Stenocactus should lose its status as genus and be included to the Ferocacti (the DNA tests above suggest otherwise
Stenocactus has long been considered closely related to Ferocactus, just as Ferocactus has long been considered a slightly dodgy genus, probably not monophyletic. The DNA supports both opinions, although most data suggests that Stenocactus can stand along as a monophyletic group. Look at the position of Ferocactus histrix in Butterworth's cladograms though, the relationships between these species are not completely clear yet. Again a hit-you-in-the-face feature like many very narrow ribs has created a genus when other features like flowers and fruit suggest they are very similar. Look at S. coptonogonus and its pretty hard to separate from a Ferocactus.